An introduction to DataLad for the ABCD ReproNim course week 8b¶
Welcome, ABCD-ReproNim students! This section belongs to week 8b of the ABCD-ReproNim course on DataLad, and contains code-along snippets to copy and paste into your own terminal, as well as additional references to useful chapters if you want to read up about specific commands or concepts elsewhere in the DataLad Handbook.
Introduction & set-up¶
In order to code along, you should have a recent DataLad version, e.g., 0.13.6 or higher, installed, and you should have a configured Git identity.
If you need them, installation, updating, and configuration instructions are in the section Installation and configuration.
If you are unsure about your version of DataLad, you can check it using the following command:
datalad --version
If you are unsure if you have configured your Git identity already, you can check if your name and email are printed to the terminal when you run
git config --get user.name
git config --get user.email
If nothing is returned, you need to configure your Git identity.
How to use DataLad¶
DataLad is a command line tool and it has a Python API. Whenever used, it is thus operated it in your terminal using the command line (as done above), or used it in scripts such as shell scripts, Python scripts, Jupyter Notebooks, and so forth. This is how you would import DataLad’s Python API:
ipython # if not installed, use python
>>> import datalad.api as dl
>>> dl.create(path='mydataset')
>>> exit
In scripts using other programming languages, DataLad commands can be invoked via system calls. Here is an example with R:
R # or use in RStudio
> system("datalad create mydataset")
DataLad datasets¶
Everything we’re doing happens in or involves DataLad datasets.
Creating a dataset from scratch is done with the datalad create command.
How can I turn an existing directory into a dataset?
By navigating into the dataset, and running datalad create -f . (manual) (with the -f/--force option).
You can take a look into the section Transitioning existing projects into DataLad on more info on how to transform existing directories into DataLad datasets.
It is advised, though, to first learn a bit of DataLad Basics first, so stay tuned.
datalad create only needs a name, and it will subsequently create a new directory under this name and instruct DataLad to manage it.
Here, the command also has an additional option, the -c yoda option.
With the -c option, datasets can be configured in a certain way at the time of creation, and yoda is a so-called run procedure.
You can find out about the details of the yoda procedure in the datalad handbook in sections Configurations to go, but in general this configuration is a very useful standard configuration for datasets for data analysis, as it preconfigures a dataset according to the yoda princples:
datalad create -c yoda myanalysis
After creating it, the dataset is a new directory, and you can “change directories” (cd) inside it:
cd myanalysis
You can take a look into the directory and file hierarchy in the dataset with the Unix tree command:
tree # lists the file structure
If you are on Windows, tree may not display individual files (on Windows, Unix commands are not always available, and sometimes, identically named commands behave differently between Unix and Windows systems). In this case, you can take a look by running explorer . to open up the file explorer.
The YODA procedure pre-created a useful directory structure and added some placeholder README files.
If you list all of the hidden files with ls -a as well, you can see that tools such as Git and DataLad operate in the background, with hidden directories and files:
ls -a # show also hidden files
Version control¶
Version controlling a file means to record its changes over time, associate those changes with an author, date, and identifier, creating a lineage of file content, and being able to revert changes or restore previous file versions.
DataLad datasets use two established version control tools: Git and git-annex.
Thanks to those tools, datasets can version control their contents, regardless of size.
Let’s see what happens when we delete placeholders in the README:
echo " " >| README.md # this overwrites existing contents
echo " " >| code/README.md
datalad status (manual) can report on the state of a dataset.
As we modified version controlled files, these files show up as being “modified” if you run it:
datalad status
What has changed compared to the files last known version state?
The git diff (manual) can tell us:
git diff
In order to save a modification one needs to use the datalad save (manual) command.
datalad save will save the current status of your dataset: It will save both modifications to known files and yet untracked files.
The -m/--message option lets you attach a concise summary of your change.
Such a commit message makes it easier for others and your later self to understand a dataset’s history:
datalad save -m "Replace placeholder in README"
Note that datalad save will save all modifications in a dataset at once!
If you have several modified files, you can supply a path to the file or files you want to save.
To demonstrate this, we make two unrelated changes: adding a new file (a comic downloaded from the web via wget), and giving the project a title:
wget https://imgs.xkcd.com/comics/compiling.png
Windows users may not have wget
If the wget command above fails for you, you could
Install a Windows version of wget
Use the following
curlcommand:curl https://imgs.xkcd.com/comics/compiling.png --output compiling.png(recent Windows builds includecurlnatively)Download and save the image from your web browser
Here’s a project title that we echo into the README:
echo "#My first data analysis with DataLad" > README.md'
With these changes, there are two modifications in your dataset, a modified file and an untracked file:
datalad status
You can add a path to make sure only modifications in the specified file are saved:
datalad save -m "Add project information to README" README.md
And perform a second datalad save to save remaining changes, i.e., the yet untracked comic:
datalad save -m "Add a motivational webcomic"
Your dataset has now started to grow a log of everything that was done.
You can view this history with the command git log (manual), or any tool that can display Git history, such as tig.
You can even ask a specific file what has been done to it:
git log README.md
While you can add and save any file into your dataset, it is often useful to know where files came from.
If you add a file from a web-source into your dataset, you can use the command datalad download-url in order to download the file, save it together with a commit message into the dataset, and record its origin internally.
Soon it will become clear why this is a useful feature.
Here, we add a comic as a little Easter egg (because we save it as a hidden dotfile called .easteregg) into the dataset:
datalad download-url -m "add motivational comic to my dataset" \
-O .easteregg.png \
https://imgs.xkcd.com/comics/fuck_grapefruit.png
# open the comic
The very first chapter of the handbook, DataLad datasets will show you even more details about version controlling files in datasets.
Data consumption & transport¶
Datasets can be installed from local paths or remote URLs using datalad clone (manual).
Cloning is a fast operation, and the resulting dataset typically takes up only a fraction of the total size of the data that it tracks:
cd ../
datalad clone git@github.com:psychoinformatics-de/studyforrest-data-phase2.git
What we have cloned is the studyforrest dataset, a neuroimaging dataset with a few Gigabytes of data. After installations, the directory tree can be browsed, but most files in datasets will not yet contain file content. This makes cloning fast and datasets lightweight:
cd studyforrest-data-phase2
ls
# print the size of the directory in human readable sizes
du -sh
How large can it get actually?
Cloned datasets can have a lot of file contents.
datalad status can report on how much data actually is accessible with the --annex and --annex all options:
datalad status --annex
On demand, content for files, directories, or the complete dataset can be downloaded using datalad get (manual).
The snippet below uses globbing to get the content of all nifti files for a localization task of one subject, but you could also get a full directory, a single file, all files, etc.:
datalad get sub-01/ses-localizer/func/sub-01_ses-localizer_task-objectcategories_run-*.nii.gz
This works because DataLad datasets track where file contents are available from.
If the origin of a file (such as a web source) is known, you can drop file content to free up disk space, but you retain access via datalad get:
datalad drop sub-01/ses-localizer/func/sub-01_ses-localizer_task-objectcategories_run-4_bold.nii.gz
This, too, works for files saved with datalad download-url (manual):
cd ../myanalysis
datalad drop .easteregg.png
but DataLad will refuse to drop files that it doesn’t know how to reobtain unless you use --nocheck:
datalad drop compiling.png
Afterward dropping files, only “metadata” about file content and file availability stays behind, and you can’t open the file anymore:
# on Windows, use "start" instead of "xdg-open"
xdg-open .easteregg.png # it is gone :(!
But because the origin of the file is known, it can be reobtained using the datalad get:
datalad get .easteregg.png
Opening the comic works again, afterwards:
# on Windows, use "start" instead of "xdg-open"
xdg-open .easteregg.png
This mechanism gives you access to data without the necessity to store all of the data locally.
As long as there is one location that data is available from (a dataset on a shared cluster, a web source, cloud storage, a USB-stick, …) and this source is known, there is no need for storing data when it is not in use.
If you want to try it with large amounts of data, checkout datasets.datalad.org, a collection of more than 200TB of open data (also called The DataLad superdataset /// because it is a dataset hierarchy that includes a large range of public datasets and can be obtained by running datalad clone \\\).
Dataset nesting¶
Datasets can be nested in superdataset-subdataset hierarchies.
This overcomes scaling issues. Some dataset that we work with including ABCD become incredibly large, and when they exceed a few 100k files version control tools can struggle and break. By nesting datasets, and you will see concrete examples later, you can overcome this and split a dataset into manageable pieces. If you are interested in finding out more, take a look into the use case Scaling up: Managing 80TB and 15 million files from the HCP release or the chapter Go big or go home.
But it also helps to link datasets as modular units together, and maximizes the potential for reuse of the individual datasets. In the context of data analysis, it is especially helpful to do this to link input data to an analysis dataset – it helps to reuse data in multiple analysis, to link input data in a precise version, and to create an intuitively structured dataset layout.
We will start a data analysis in the myanalysis dataset.
First, let’s install input data (a small dataset from GitHub) as a subdataset.
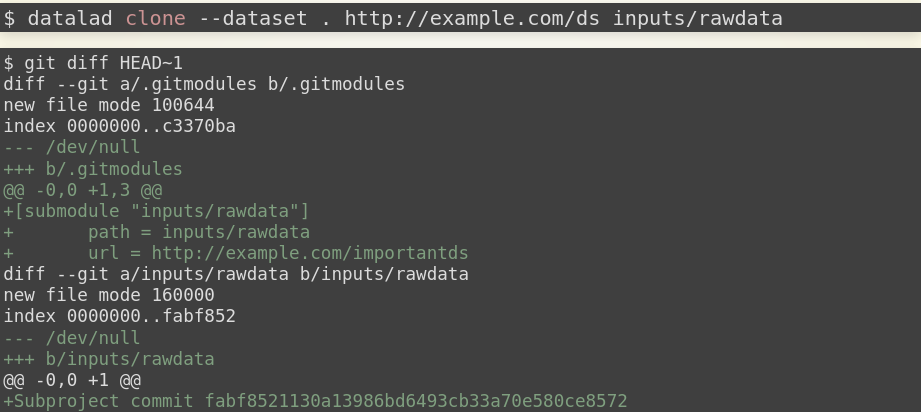
This is done with the -d/--dataset option of datalad clone:
datalad clone -d . git@github.com:datalad-handbook/iris_data.git input/
This dataset has been linked in a precise version to the dataset, and it has preserved its complete history (if you are on a native Windows installation, please run git show main instead – the reason for this is explained in the first chapter of the handbook):
# this shows details of the last entry in your dataset history
git show
Navigate into the subdataset and see for yourself that it has a standalone history, and that its most recent commit shasum is identical to the subproject commit that is registered in the superdataset:
cd input
git log
The YODA principles¶
The YODA principles are guidelines on the structure, content, and handling of data analyses. They aren’t limited to DataLad, but they can be easily adopted if you’re using DataLad. You can find a complete section on them, including the upcoming data analysis example and a section on how to work with computational environments starting from the section YODA: Best practices for data analyses in a dataset.
Reproducible analyses¶
Not only can DataLad version control, consume, and share data, it can also help to create datasets with data analyses in a way that your future self and others can easily and automatically recompute what was done. In this part of the tutorial, we start with a small analysis to introduce core commands and concepts for reproducible execution.
For this small analysis, we start by adding some code for a data analysis (copy paste from cat to the final EOT to paste the code into a file scripty.py in your code/ directory, or use an editor of your choice and copy paste the script):
cat << EOT > code/script.py
import pandas as pd
import seaborn as sns
import datalad.api as dl
from sklearn import model_selection
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import classification_report
data = "input/iris.csv"
# make sure that the data are obtained (get will also install linked sub-ds!):
dl.get(data)
# prepare the data as a pandas dataframe
df = pd.read_csv(data)
attributes = ["sepal_length", "sepal_width", "petal_length","petal_width", "class"]
df.columns = attributes
# create a pairplot to plot pairwise relationships in the dataset
plot = sns.pairplot(df, hue='class', palette='muted')
plot.savefig('pairwise_relationships.png')
# perform a K-nearest-neighbours classification with scikit-learn
# Step 1: split data in test and training dataset (20:80)
array = df.values
X = array[:,0:4]
Y = array[:,4]
test_size = 0.20
seed = 7
X_train, X_test, Y_train, Y_test = model_selection.train_test_split(X, Y,
test_size=test_size,
random_state=seed)
# Step 2: Fit the model and make predictions on the test dataset
knn = KNeighborsClassifier()
knn.fit(X_train, Y_train)
predictions = knn.predict(X_test)
# Step 3: Save the classification report
report = classification_report(Y_test, predictions, output_dict=True)
df_report = pd.DataFrame(report).transpose().to_csv('prediction_report.csv')
EOT
This script highlights an important key point from the YODA principles: relative paths instead of absolute paths make the dataset self-contained and portable. Results are saved into the top level dataset, and not next to the input data. It also demonstrates how DataLad’s Python API can be used with a dl.get() function in the script.
Running the above code block created a new file in the dataset:
datalad status
Let’s save it with a datalad save command.
DataLad save can in addition also attach an identifier in the form of a tag with the --version-tag flag:
datalad save -m "add script for kNN classification and plotting" \
--version-tag ready4analysis code/script.py
The datalad run (manual) command can run this script in a way that links the script to the results it produces and the data it was computed from.
In principle, the command is simple: Execute any command, save the resulting changes in the dataset, and associate them as well as all other optional information provided.
Because each datalad run ends with a datalad save, it is recommended to start with a clean dataset (see DataLad, run! for details on how to use it in unclean datasets):
datalad status
Then, give the command you would execute to datalad run, in this case python code/script.py.
Datalad will take the command, run it, and save all of the changes in the dataset that this leads this to under the commit message specified with the -m option.
Thus, it associates the script (or any command execution) with the results it generates.
But the command can become even more helpful.
Below, we also specify the input data the command needs - DataLad will make sure to datalad get the data beforehand.
And we also specify the output of the command.
This is not in order to identify outputs (DataLad would do that on its own), but to specify files that should be datalad unlock (manual)ed and potentially updated if the command is reran – but more on this later.
To understand fully what --output does, please read chapters DataLad, run! and Under the hood: git-annex:
datalad run -m "analyze iris data with classification analysis" \
--input "input/iris.csv" \
--output "prediction_report.csv" \
--output "pairwise_relationships.png" \
"python3 code/script.py"
software note
In order to execute the above script successfully you will need to run it in an environment that has the Python packages pandas, scikit-learn, datalad, and seaborn installed. If you’re thinking “WTF, it is SO inconvenient that I have to create the software environment to make this run”, wait until the next section.
Datalad creates a commit in the dataset history. This commit has the commit message as a human readable summary of what was done, it contains the produced output, and it has a machine readable record that contains information on the input data, the results, and the command that was run to create this result:
# take a look at the most recent entry in git log
git log -n 1
This machine readable record is particularly helpful, because one can now instruct datalad to rerun this command so that you don’t have to memorize what had been done, and people you share the dataset with don’t need to ask you how this result was produced, by can simply let DataLad tell them.
This is done with the datalad rerun command.
For this demonstration, there is a published analysis dataset that resembles the one created here fully at github.com/adswa/my_analysis.
This dataset can be cloned, and the analysis within it can be automatically rerun:
cd ../
datalad clone git@github.com:adswa/myanalysis.git analysis_clone
Among other ways, run records can be identified via their commit hash.
If given to datalad rerun <hash>, DataLad will read the machine readable record of what was done, get required data, unlock to-be-modified files, and recompute the exact same thing:
cd analysis_clone
git log pairwise_relationships.png
# this is the start of commit hash of the run record
datalad rerun 71cb8c5
This allows others to very easily rerun computations, but it also spares yourself the need to remember how a script was executed, and results can simply be asked where they came from.
Computational reproducibility¶
Its fantastic to have means to recompute a command automatically, but the ability to re-execute a command is often not enough. If you don’t have the required Python packages available, or in a wrong version, running the script and computing the results will fail. In order to be computationally reproducible the run record does not only need to link code, command, and data, but also encapsulate the software that is necessary for a computation:
cd ../myanalysis
The way this can be done is with a DataLad extension called datalad container.
You can install this extension with pip by running pip install datalad-container.
This extension allows to attach software containers such as Singularity or Docker container images to the dataset and execute commands inside of these containers.
Thus, the dataset can share share data, code, code execution, and software.
Here is how this works: First, attach a software container to the dataset using datalad containers-add.
This command needs a name for the container (here it is called software, but you can go for any name – how about “take-this-one-mom”?), and a URL or path where to find the container.
Here, it is a URL that points to Singularity-hub (but Docker-Hub, with a docker://<user>/<container>:<version> URL, would work fine, too).
This records a pre-created software environment with the required Python packages in the dataset:
datalad containers-add software --url shub://adswa/resources:2
Software note
You need to have a software that can work with software containers – either singularity or Docker!
Why may Singularity be a better choice than Docker?
Singularity, unlike Docker, can be deployed on shared compute infrastructure such as computational clusters as it does not require or grant superuser privileges (“sudo rights”) to users that use a container. Docker is not deployed on HPC systems is because it grants users those sudo rights, and on multi-user systems users should not have those privileges, as it would enable them to tamper with other’s or shared data and resources, posing a severe security threat. Singularity is capable of working with both Docker and Singularity containers, though.
Afterwards, rerun the analysis in the software container with the datalad containers-run command.
This container works just as the run command before, with the additional -n/--name option that is needed to specify the container name.
DataLad then executes this command inside of the container image, and if you were to rerun such an analysis, DataLad would not only retrieve the input data but also the software container:
datalad containers-run -m "rerun analysis in container" \
--container-name software \
--input "input/iris.csv" \
--output "prediction_report.csv" \
--output "pairwise_relationships.png" \
"python3 code/script.py"
You can read more about this command and containers in general in the section Computational reproducibility with software containers.
The ABCD data as a dataset¶
At the time that the lecture is recorded, retrieving ABCD data is not yet possible with DataLad and needs to be done via NDA Python tools or its web interface.
What is difficult for us about turning the data into a DataLad dataset is that it contains filenames with GUIDs, and those can’t be shared publicly. However, we’re working on a solution that would enable you to clone the ABCD data easily, using NDA credentials with appropriate access.
This section gives a sneak peek into how an ABCD DataLad dataset feels like. Because the ABCD dataset is super large, it is split into a hierarchy of nested datasets. There is one superdataset (the one that everyone would clone), and this superdataset contains one subdataset per participant, and each participant dataset can also contain additional subdatasets.
This splits the vast amount of files in the ABCD data between thousands of datasets.
When working with a nested hierarchy of datasets, the subdatasets aren’t installed automatically when you install the top-level dataset with datalad clone.
Uninstalled datasets look like empty directories on first sight – you will not be able to browse through their file hierarchy until the are installed.
In order to install a subdataset, run datalad get.
To not automatically download data, append the -n/--no-data flag.
If you want to install all subdatasets, run datalad get -n -r . in the superdataset to install all subdatasets recursively.
If you want to practice or get a feel for datasets of this size, you can try cloning github.com/datalad-datasets/human-connectome-project-openaccess, the complete human connectome project data. Useful tips for working with large dataset hierarchies are in the section Gists.